News

From text to traceability: RRID integration in Europe PMC
Hinxton, UK & San Diego, US — 11 May 2026
Scientific articles are theoretically rich in details about antibodies, cell lines, model organisms, or software tools. However, these details are often poorly reported and remain locked in unstructured text, making it difficult, or impossible, to reproduce experiments. Mentions are often ambiguous, inconsistent, or difficult to track across studies. This is where Research Resource Identifiers (RRIDs) come to the rescue. RRIDs are unique persistent identifiers (PIDs) assigned to key research resources and are structured (for example, RRID:AB_572263) to transform a vague mention (like “anti-GFP antibody”) into something precise, traceable, and comparable across studies. Unlike plain text mentions, these PIDs are globally unique, machine-readable, and linked to curated metadata.
How RRIDs are maintained and developed
RRIDs are developed and maintained by the FAIR Data Informatics laboratory at the University of California at San Diego and RRIDs.org (a 501(c)(3) not-for-profit organization) is the organization that owns RRIDs. The RRID platform aggregates and curates biomedical resource information and links to usage of RRIDs in the scientific literature to create a structured, traceable connection between a resource and its documented use. Correctly reported resources are easier to find, but sometimes resources are mentioned but not properly identified, are ambiguous, or even incorrect. SciCrunch has developed machine learning methods to identify these harder-to-find connections. This enrichment layer enables downstream platforms like Europe PMC to surface RRIDs even when authors did not explicitly structure them in the article text, the supplemental files, or reagent tables. These previously opaque references can, therefore, become discoverable, queryable, and reusable, which is an important step toward more transparent and reproducible research.
How to discover RRIDs in Europe PMC
RRIDs are integrated into Europe PMC through the annotation platform. RRIDs provided by SciCrunch are associated with articles within the Europe PMC database, making them available through search, integrated into the SciLite annotations tool, and retrievable programmatically through the Annotations API. API access allows developers to retrieve and use RRID annotations linked to literature in Europe PMC in their own applications and researchers retrieve the data for analysis.
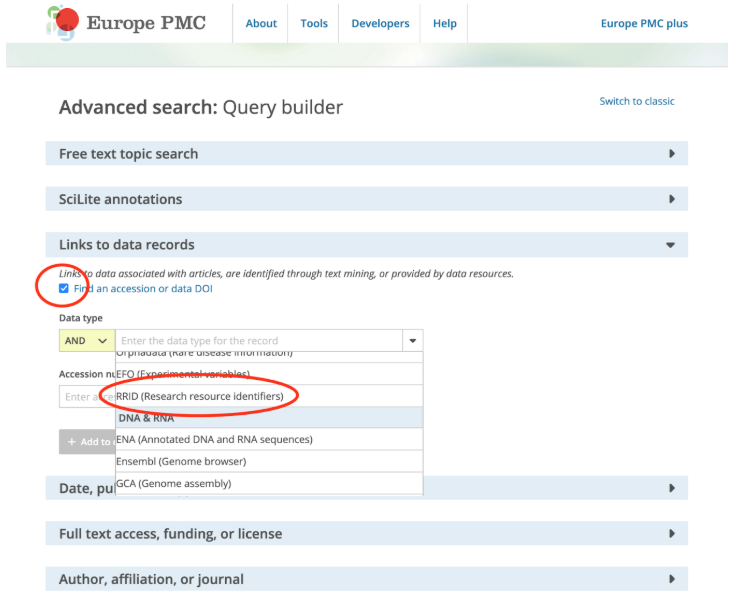
RRIDs can be searched within Europe PMC through the Advanced search tool. In the ‘Links to data records’ section, tick the ‘Find an accession or data DOI’ checkbox and then select the ‘RRID (Research resource identifiers)’ under the ‘Data type’ drop-down.
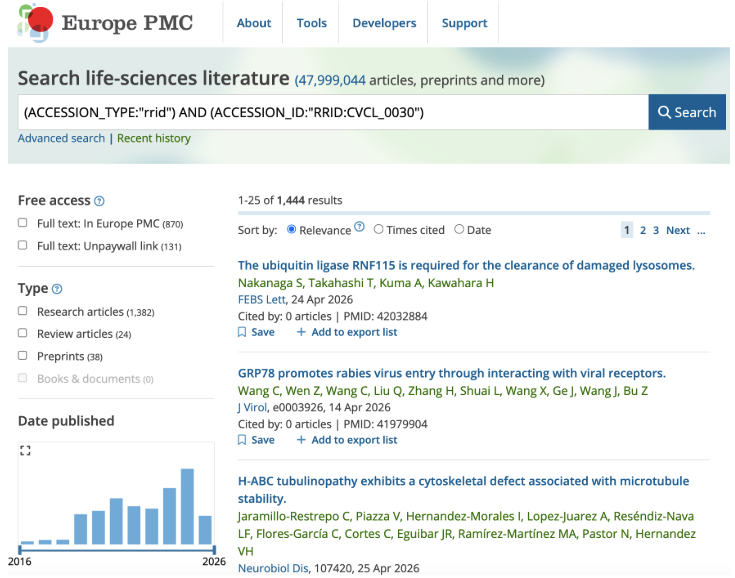
You may then enter an RRID in the “Accession number or data DOI” box and add this to your search. For example, the RRID for HeLa cells is RRID:CVCL_0030. Searching for this using Advanced search produces the following results.
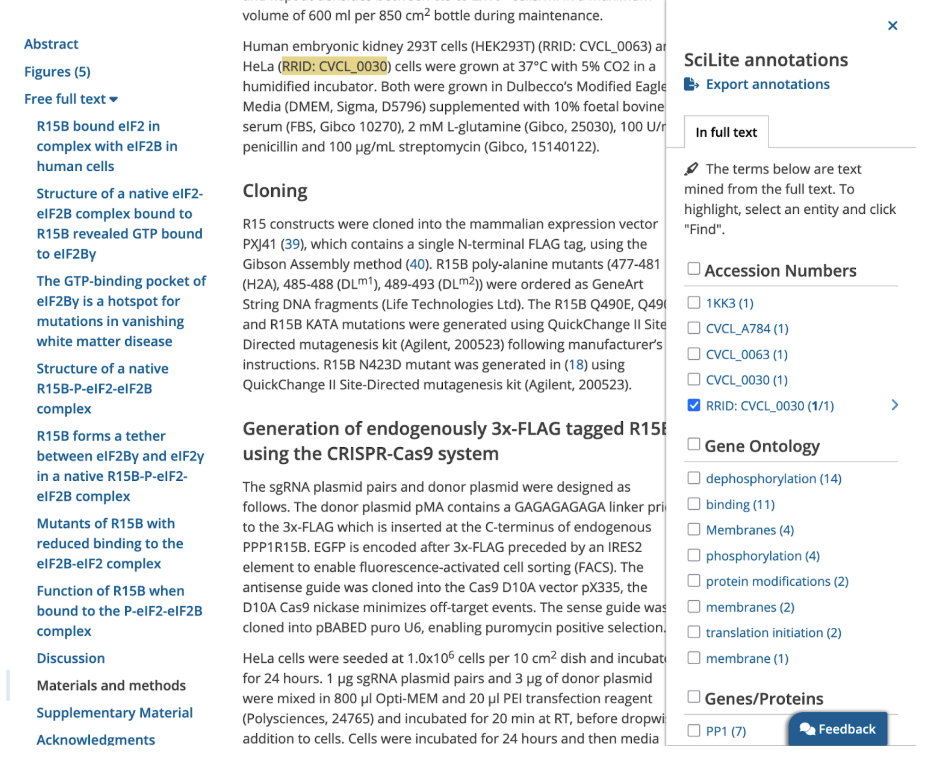
Reading an article, you can find references to RRIDs using the SciLite tool.
Benefits to users of Europe PMC
Through this collaboration, Europe PMC now indexes over one million RRIDs (in over a hundred thousand papers), transforming previously fragmented data into a structured, discoverable dataset of literature and associated RRIDs combined. This integration enhances reproducibility, supports large-scale meta-analyses, and enables downstream reuse in emerging tools and workflows. By combining RRIDs with other search terms, users can perform precise, targeted queries, significantly improving the specificity and utility of searches across the Europe PMC platform.
As RRID coverage and adoption continue to grow, these connections could support new approaches to reproducibility monitoring, resource validation, trend analysis, and AI-driven discovery tools built on structured scientific knowledge rather than unstructured text alone.
Written by Matthew Jeffryes and Melissa Harrison from Europe PMC and Anita Bandrowski and Martijn Roelandse from SciCrunch.